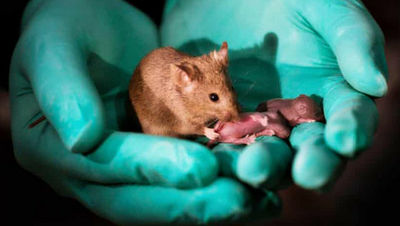
A report indicates that changing the DNA sequence of female mice by just one letter resulted in the development of male reproductive organs.
A research team at Bar-Ilan University in Israel has reported that by making only slight changes to specific non-coding DNA regions in female mouse embryos, individuals with XX chromosomes, which are normally female, developed male reproductive organs and testes. This demonstrates that extremely small changes in the mouse DNA sequence, such as the insertion of just one letter or the deletion of three letters, can trigger a dramatic developmental shift in the embryo's sexual differentiation.
A single-nucleotide enhancer mutation overrides chromosomal sex to drive XX male development | Nature Communications
Changing 'just one DNA letter' in female mice triggers growth of male genitalia | Live Science
https://www.livescience.com/health/genetics/changing-just-one-dna-letter-in-female-mice-triggers-growth-of-male-genitalia
In mammals, sex is basically determined by the SRY gene on the Y chromosome, which activates the SOX9 gene, initiating testicular development. SOX9 is a 'master gene' essential for testicular formation, and whether or not it is activated determines whether an individual becomes male or female.
In 2018, the research team identified a DNA sequence called 'Enh13' that functions as a switch for the SOX9 gene. Enh13 is a non-coding region that does not have a blueprint for the protein, but it is located upstream of SOX9 and strictly controls its expression.
Previous studies have shown that completely removing Enh13 from male embryos with XY chromosomes prevents SOX9 activation, resulting in the individual developing as a female. In this study, the opposite was done: the Enh13 sequence was slightly edited in female embryos with XX chromosomes.
The research team performed a procedure to insert one base or delete three bases at a specific site within the Enh13 molecule. When this procedure was applied to the Enh13 molecule present in both pairs of XX chromosomes, mice with XX chromosomes exhibited fully male-like external genitalia and internally developed testes. On the other hand, XX mice in which only one chromosome of the Enh13 molecule was mutated showed normal ovaries.
Below are, from left to right, a female mouse, a female mouse with a single base inserted into Enh13 of the XX chromosome, and a male mouse. All are 6-week-old adults, with the top row showing their external appearance and the bottom row showing their internal genitalia. The gene-edited mouse in the center has XX chromosomes, yet it has small testes. According to the research team, this mouse also had male external genitalia.
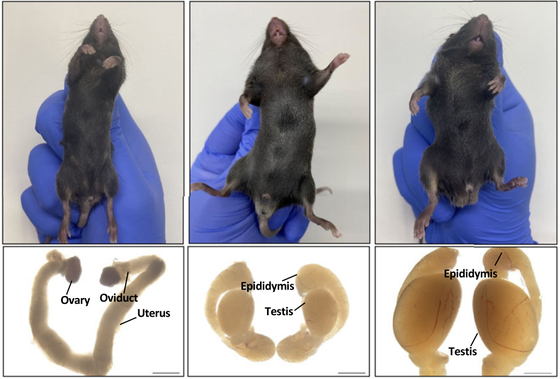
Analysis revealed that the edited Enh13 gene, despite the absence of the SRY gene, had acquired the ability to activate the SOX9 gene. However, these XX male individuals lacked a Y chromosome, and therefore lacked sperm-producing cells in their testes, resulting in their inability to reproduce.
Normally, in female mouse embryos, the activation of the SOX9 gene needs to be strongly suppressed, and it has been shown that
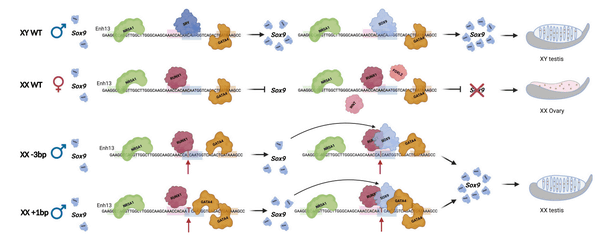
SOX9 has a self-looping property that, once activated, further amplifies its own activity, which is thought to ultimately lead to a strong activation and complete the male development program. The research team argued that this proves that even slight changes in non-coding regions can completely disrupt the balance of a complex gene network.
This discovery suggests that not only the 'blueprint' of genes themselves, but also slight abnormalities in the 'regulatory switches' that control them, can cause disorders of sex development in humans. The research team says that these findings may be useful in identifying the etiologies of previously unknown diseases and are important for understanding the importance of mutations in non-protein-coding DNA regions.
Related Posts:
in Science, Posted by log1i_yk